Thanks for the explanation! Please take all the time you need!
×
INDI Library v2.0.7 is Released (01 Apr 2024)
Bi-monthly release with minor bug fixes and improvements
Fits Viewer: Median computation wrong?
- Peter Sütterlin
-
 Topic Author
Topic Author
- Offline
- Supernova Explorer
-

- Posts: 1009
- Thank you received: 133
Fits Viewer: Median computation wrong? was created by Peter Sütterlin
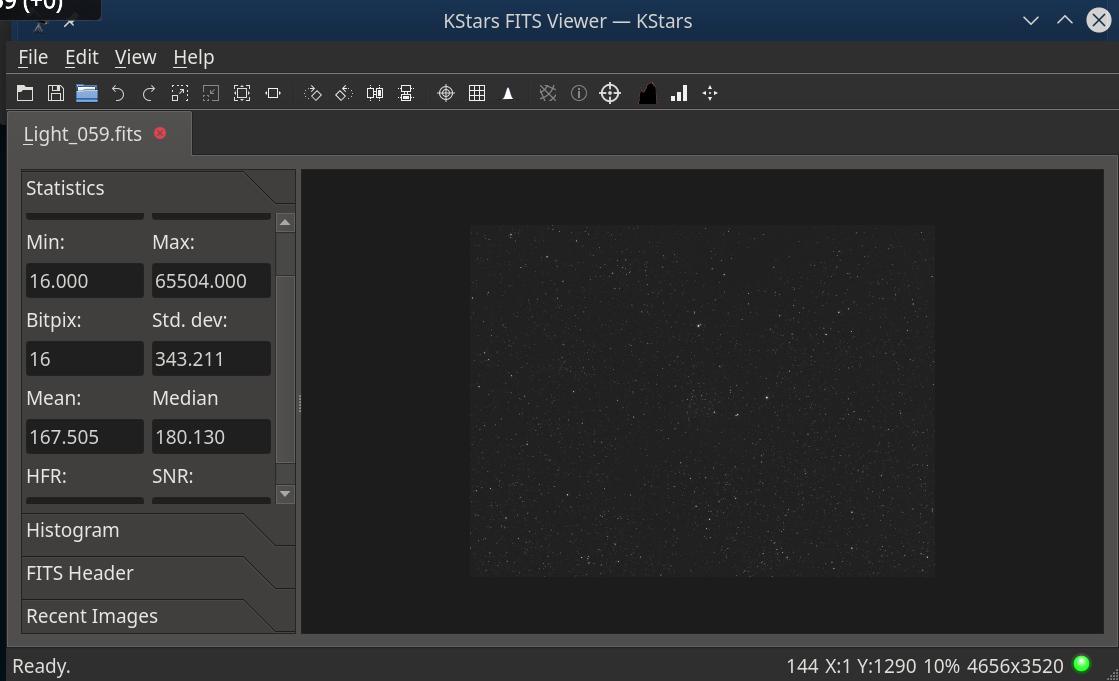
Looking at the statistics of my acquired image I was puzzled by the value for Median being higher than the average:
180.13 vs. 167.505. Reading the image in IDL and computing average, median and standard deviation I get similar values for avg and stdev, but median is 160, lower than the average (as I'd expect for such data).
180.13 vs. 167.505. Reading the image in IDL and computing average, median and standard deviation I get similar values for avg and stdev, but median is 160, lower than the average (as I'd expect for such data).
4 years 7 months ago
#42731
Please Log in or Create an account to join the conversation.
- Jasem Mutlaq
-

- Online
- Administrator
-

Replied by Jasem Mutlaq on topic Fits Viewer: Median computation wrong?
I suspect it might be due to this: cgit.kde.org/kstars.git/commit/?id=151f4...972afa331301e73886ae
can you verify?
can you verify?
4 years 7 months ago
#42735
Please Log in or Create an account to join the conversation.
- Peter Sütterlin
-
 Topic Author
Topic Author
- Offline
- Supernova Explorer
-

- Posts: 1009
- Thank you received: 133
Replied by Peter Sütterlin on topic Fits Viewer: Median computation wrong?
So I reverted this patch and recompiled. Thought I can check it by just loading that example file again, but it seems median is not computed for loaded data, only when it arrives from the camera?
So I had to test/compare using skyflat or dark data. But for those I cannot see a substantial difference between the two versions and my external computation of the median. So no real answer for now. Could of course be that only working on a (random?) subset of the data, while safe for more homogeneous images, is a bad choice for data like starfields?
So I had to test/compare using skyflat or dark data. But for those I cannot see a substantial difference between the two versions and my external computation of the median. So no real answer for now. Could of course be that only working on a (random?) subset of the data, while safe for more homogeneous images, is a bad choice for data like starfields?
4 years 7 months ago
#42746
Please Log in or Create an account to join the conversation.
- Jasem Mutlaq
-

- Online
- Administrator
-

Replied by Jasem Mutlaq on topic Fits Viewer: Median computation wrong?
Try to view the histogram and see if that makes it recompute it.
4 years 7 months ago
#42747
Please Log in or Create an account to join the conversation.
- Peter Sütterlin
-
 Topic Author
Topic Author
- Offline
- Supernova Explorer
-

- Posts: 1009
- Thank you received: 133
Replied by Peter Sütterlin on topic Fits Viewer: Median computation wrong?
No, had tried that, as the commit hinted to being related to histogram stuff.
Ah no, wait. The histogram does have its own median field, that contains a value. Had overlooked that.
But I do get the same (high) value of 180 for the median with and without that commit.
Playing around with it a bit more, median really behaves strange. It changes substantially when the image is clipped. E.g., load an image, activate histogram. I get mean 167.5, median 180.1. The limit slider (only R, it's a monochrome image) is at 16 and 65504. Change the lower clip to 17 and click 'Apply', and median changes to 181.1! mean stays unchanged. Clipping an array will never change the median, unless the low clip is higher than the unclipped median (or the high one lower). Also, for an (u)int array, it should not be a fractional number.
Ah no, wait. The histogram does have its own median field, that contains a value. Had overlooked that.
But I do get the same (high) value of 180 for the median with and without that commit.
Playing around with it a bit more, median really behaves strange. It changes substantially when the image is clipped. E.g., load an image, activate histogram. I get mean 167.5, median 180.1. The limit slider (only R, it's a monochrome image) is at 16 and 65504. Change the lower clip to 17 and click 'Apply', and median changes to 181.1! mean stays unchanged. Clipping an array will never change the median, unless the low clip is higher than the unclipped median (or the high one lower). Also, for an (u)int array, it should not be a fractional number.
IDL> help,p
P UINT = Array[4656, 3520]
IDL> print,avg(p),avg(p>20000),avg(p>20000<22500)
22400.446 22402.729 22186.562
IDL> print,median(p),median(p>20000),median(p>20000<22500)
22480.0 22480.0 22480.0
4 years 7 months ago
#42750
Please Log in or Create an account to join the conversation.
- Hy Murveit
-

- Away
- Administrator
-

- Posts: 1223
- Thank you received: 566
Replied by Hy Murveit on topic Fits Viewer: Median computation wrong?
Jasem and Peter,
I looked into this briefly, and saw that the median computation is not an exact one. That is, the code in fitshistogram.cpp divides the data value's range into at most 400 bins, counts how many sample values fall into each bin, finds which bin contains the median, and then uses the lowest range from the bin as the median. (An exact computation would need to resample the data, and count each possible sample value that falls into the 'median bin'). Note, btw, that the median should be an integer, and in Peter's example it is 180.1.
If the sample data ranges from 0-64K, and there are 400 bins, then the bin size is 163, so the median value could be off by that much. If its 0->4K, then the error could be 10.
See around line 270 in fitshistogram.cpp:
median[n] = i * binWidth[n] + FITSMin[n];
This is likely the issue Peter is bringing up.
Since I plan on working some on that file, I'd be happy to look into this, if you like Jasem, but it may be several weeks until I get to it.
Hy
I looked into this briefly, and saw that the median computation is not an exact one. That is, the code in fitshistogram.cpp divides the data value's range into at most 400 bins, counts how many sample values fall into each bin, finds which bin contains the median, and then uses the lowest range from the bin as the median. (An exact computation would need to resample the data, and count each possible sample value that falls into the 'median bin'). Note, btw, that the median should be an integer, and in Peter's example it is 180.1.
If the sample data ranges from 0-64K, and there are 400 bins, then the bin size is 163, so the median value could be off by that much. If its 0->4K, then the error could be 10.
See around line 270 in fitshistogram.cpp:
median[n] = i * binWidth[n] + FITSMin[n];
This is likely the issue Peter is bringing up.
Since I plan on working some on that file, I'd be happy to look into this, if you like Jasem, but it may be several weeks until I get to it.
Hy
The following user(s) said Thank You: Jasem Mutlaq, Peter Sütterlin
4 years 7 months ago
#42765
Please Log in or Create an account to join the conversation.
- Peter Sütterlin
-
 Topic Author
Topic Author
- Offline
- Supernova Explorer
-

- Posts: 1009
- Thank you received: 133
Replied by Peter Sütterlin on topic Fits Viewer: Median computation wrong?
Thanks Hy for this enlightening explanation. Well received!
So for most astronomical images that would mean that most of the data will be in bin 1 or 2, and it's the computation of the bin width that determines the median value? And its value changes, because the clipping range boundaries are used to compute the width. Sounds like a nasty issue if you want to solve it for various data types.... could a variable bin width (e.g., logarithmic) help there?
So for most astronomical images that would mean that most of the data will be in bin 1 or 2, and it's the computation of the bin width that determines the median value? And its value changes, because the clipping range boundaries are used to compute the width. Sounds like a nasty issue if you want to solve it for various data types.... could a variable bin width (e.g., logarithmic) help there?
4 years 7 months ago
#42769
Please Log in or Create an account to join the conversation.
- Hy Murveit
-

- Away
- Administrator
-

- Posts: 1223
- Thank you received: 566
Replied by Hy Murveit on topic Fits Viewer: Median computation wrong?
Assuming that I got it right (and I only spent 10 minutes looking at the code) I don't think most astro data will use bin 1 or 2. A lot of folks try and expose so their median is ~500-1000 on a 64K scale, which would be bins 4-7 or so. And that assume Ekos is using the full 64K range, and it may not . In any event, I believe it is approximating in a way that can be improved with modest extra computation.
I don't think this is difficult to solve. I would keep the first pass computation as is. I would then take a 2nd pass over the sample data, which (if it used the same downsampling code as Jasem pointed to in the commit above) wouldn't be too expensive. So, if it knew the median was in bin 4 (e.g. perhaps between values 640 and 800), it would just run through all the sample data (or e.g. every 10th sample) and count the number of samples with values 640, 641, 642, ... 800, and then it could use that table to compute an exact median--well the median would be exact for the downsampled cohort, or totally exact if there was no downsampling.
Hy
I don't think this is difficult to solve. I would keep the first pass computation as is. I would then take a 2nd pass over the sample data, which (if it used the same downsampling code as Jasem pointed to in the commit above) wouldn't be too expensive. So, if it knew the median was in bin 4 (e.g. perhaps between values 640 and 800), it would just run through all the sample data (or e.g. every 10th sample) and count the number of samples with values 640, 641, 642, ... 800, and then it could use that table to compute an exact median--well the median would be exact for the downsampled cohort, or totally exact if there was no downsampling.
Hy
4 years 7 months ago
#42772
Please Log in or Create an account to join the conversation.
- Jasem Mutlaq
-

- Online
- Administrator
-

Replied by Jasem Mutlaq on topic Fits Viewer: Median computation wrong?
Thanks for the explanation! Please take all the time you need!
4 years 7 months ago
#42778
Please Log in or Create an account to join the conversation.
Time to create page: 0.534 seconds
© 2003-2022 by INDI Library. All rights reserved.